ATACdb v1.03
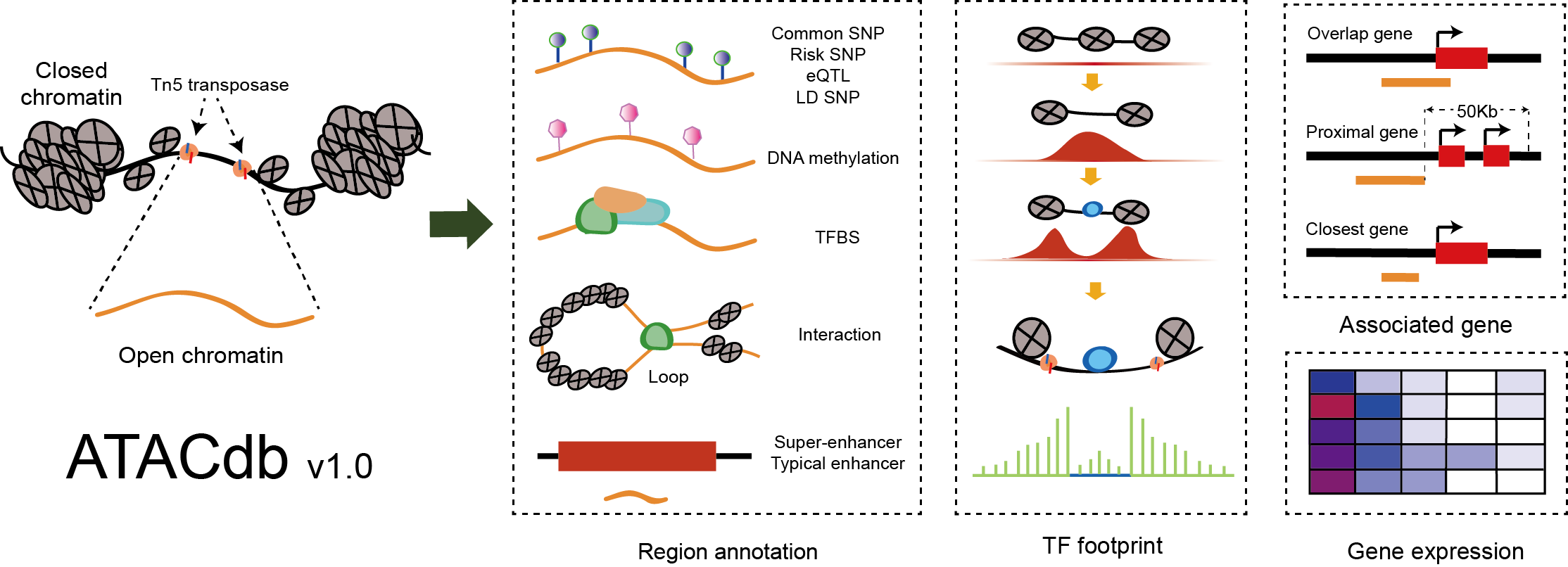
A comprehensive human chromatin accessibility database
 30/8/2020 ATACdb v1.01: Update the page of 'Home' and 'Submit'.
30/8/2020 ATACdb v1.01: Update the page of 'Home' and 'Submit'.
 06/10/2021 ATACdb v1.02: Update the download files to provide the associated genes of chromatin accessibility region.
06/10/2021 ATACdb v1.02: Update the download files to provide the associated genes of chromatin accessibility region.
 18/10/2021 ATACdb v1.03: Update the page of 'Help' and 'Download'.
18/10/2021 ATACdb v1.03: Update the page of 'Help' and 'Download'.
Wang F, Bai X, Wang Y, Jiang Y, Ai B, Zhang Y, Liu Y, Xu M, Wang Q, Han X, Pan Q, Li Y, Li X, Zhang J, Zhao J, Zhang G, Feng C, Zhu J, Li C. ATACdb: a comprehensive human chromatin accessibility database. Nucleic Acids Res. 2020 Oct 30:gkaa943. doi: 10.1093/nar/gkaa943. Epub ahead of print. PMID: 33125076.
SEdb
SEdb: the comprehensive human Super-Enhancer database
ENdb
ENdb: an experimentally supported enhancer database for human and mouse
SEanalysis
SEanalysis: a web tool for super-enhancer associated regulatory analysis
LncSEA
LncSEA: a comprehensive human lncRNA sets resource and enrichment analysis platform
KnockTF
KnockTF: a comprehensive human gene expression profile database with knockdown/knockout of transcription
factors
TRCirc
TRCirc: a resource for transcriptional regulation information of circRNAs
TRlnc
TRlnc: a comprehensive database of human transcriptional regulation of lncRNAs
VARAdb
VARAdb: a variation annotation database for human